26. Schelling Model with NumPy#
26.1. Overview#
In the previous lecture, we implemented the Schelling segregation model using pure Python and standard libraries.
In this lecture, we will rewrite the model using NumPy arrays and functions.
This is intended as a first step towards greater efficiency.
In later lectures, we’ll improve execution speed further by adopting JAX and modifying algorithms.
We’ll achieve greater speed — but at the cost of readability!
import matplotlib.pyplot as plt
import numpy as np
from numpy.random import uniform
import time
26.2. Data Representation#
In the class-based version, each agent was an object storing its own type and location.
Here we take a different approach: we store all agent data in NumPy arrays.
locations— an \(n \times 2\) array where row \(i\) holds the \((x, y)\) coordinates of agent \(i\)types— an array of length \(n\) where entry \(i\) is 0 or 1, indicating agent \(i\)’s type
Let’s set up the parameters:
num_of_type_0 = 1000 # number of agents of type 0 (orange)
num_of_type_1 = 1000 # number of agents of type 1 (green)
n = num_of_type_0 + num_of_type_1 # total number of agents
num_neighbors = 10 # number of agents viewed as neighbors
max_other_type = 6 # max number of different-type neighbors tolerated
Here’s a function to initialize the state with random locations and types:
def initialize_state():
locations = uniform(size=(n, 2))
types = np.zeros(n, dtype=int)
types[num_of_type_0:] = 1
return locations, types
Let’s see what this looks like:
np.random.seed(1234)
locations, types = initialize_state()
print(f"locations shape: {locations.shape}")
print(f"First 5 locations:\n{locations[:5]}")
print(f"\ntypes shape: {types.shape}")
print(f"First 20 types: {types[:20]}")
locations shape: (2000, 2)
First 5 locations:
[[0.19151945 0.62210877]
[0.43772774 0.78535858]
[0.77997581 0.27259261]
[0.27646426 0.80187218]
[0.95813935 0.87593263]]
types shape: (2000,)
First 20 types: [0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0]
26.3. Helper Functions#
Let’s write some functions that compute what we need while operating on the arrays.
26.3.1. Checking Happiness#
An agent is happy if at most max_other_type of their nearest neighbors
are of a different type:
def is_happy(i, locations, types):
" True if agent i has at most max_other_type neighbors of a different type. "
# Compute distance from agent i to every other agent
distances = np.linalg.norm(locations[i] - locations, axis=1)
distances[i] = np.inf # exclude self
neighbors = np.argsort(distances)[:num_neighbors] # indices of nearest
num_other = np.sum(types[neighbors] != types[i])
return num_other <= max_other_type
26.3.2. Moving Unhappy Agents#
When an agent is unhappy, they keep trying new random locations until they find one where they’re happy:
def move_agent(i, locations, types, max_attempts=10_000):
" Move agent i to a new location where they are happy. "
attempts = 0
while not is_happy(i, locations, types) and attempts < max_attempts:
locations[i, :] = uniform(), uniform()
attempts += 1
Note that locations[i, :] = ... modifies the array in place — the change
is visible to all code that references locations.
26.4. Visualization#
Here’s some code for visualization — we’ll skip the details
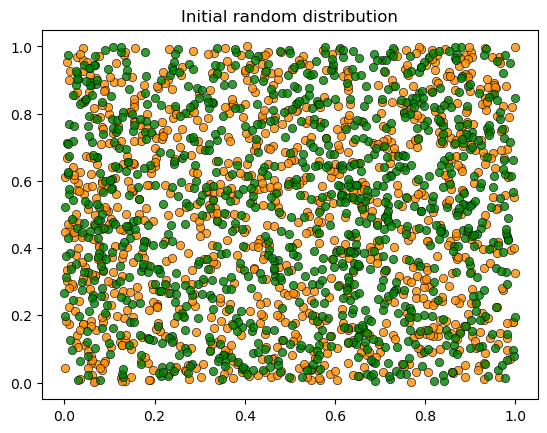
Let’s visualize the initial random distribution:
26.5. The Simulation#
Now we put it all together.
As in the first lecture, each iteration cycles through all agents in order, giving each the opportunity to move:
def run_simulation(max_iter=100_000, seed=42):
"""
Run the Schelling simulation.
Each iteration cycles through all agents, giving each a chance to move.
"""
np.random.seed(seed)
locations, types = initialize_state()
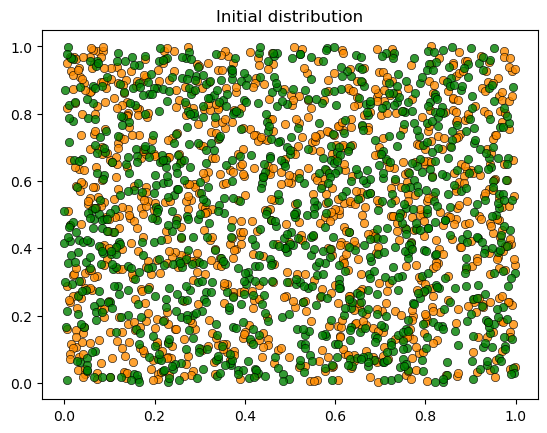
plot_distribution(locations, types, 'Initial distribution')
# Loop until no agent wishes to move
start_time = time.time()
converged = False
for iteration in range(1, max_iter + 1):
print(f'Entering iteration {iteration}')
someone_moved = False
for i in range(n):
if not is_happy(i, locations, types):
move_agent(i, locations, types)
someone_moved = True
if not someone_moved:
converged = True
break
elapsed = time.time() - start_time
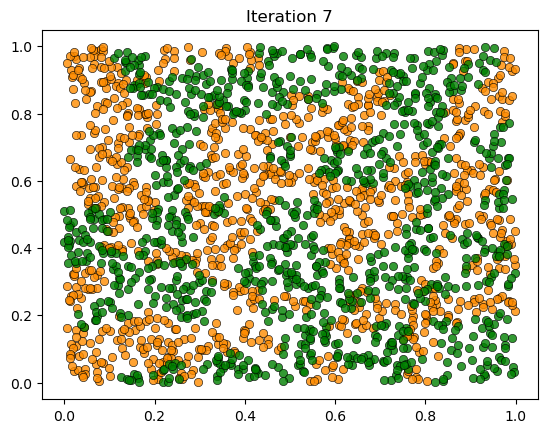
plot_distribution(locations, types, f'Iteration {iteration}')
if converged:
print(f'Converged in {elapsed:.2f} seconds after {iteration} iterations.')
else:
print('Hit iteration bound and terminated.')
return locations, types
26.6. Results#
Let’s run the simulation:
locations, types = run_simulation()
We see the same phenomenon as in the class-based version: starting from a random mixed distribution, agents self-organize into segregated clusters.
26.7. Performance#
The NumPy version is faster than the pure Python version, but still slow for large populations.
In the next lecture, we’ll rewrite the model using JAX, which offers just-in-time compilation, GPU acceleration, and faster nearest neighbor computations.